VEGA documentation
VEGA is a deep generative model for scRNA-Seq data whose decoder structure is informed by gene modules such as pathways, gene regulatory networks, or cell type marker sets. It allows embedding of single-cell data into an interpretable latent space, inference of gene module activity at the single-cell level, and differential activity testing for those gene modules between groups of cells. VEGA is implemented in Pytorch and works around the scanpy and scvi-tools ecosystems.
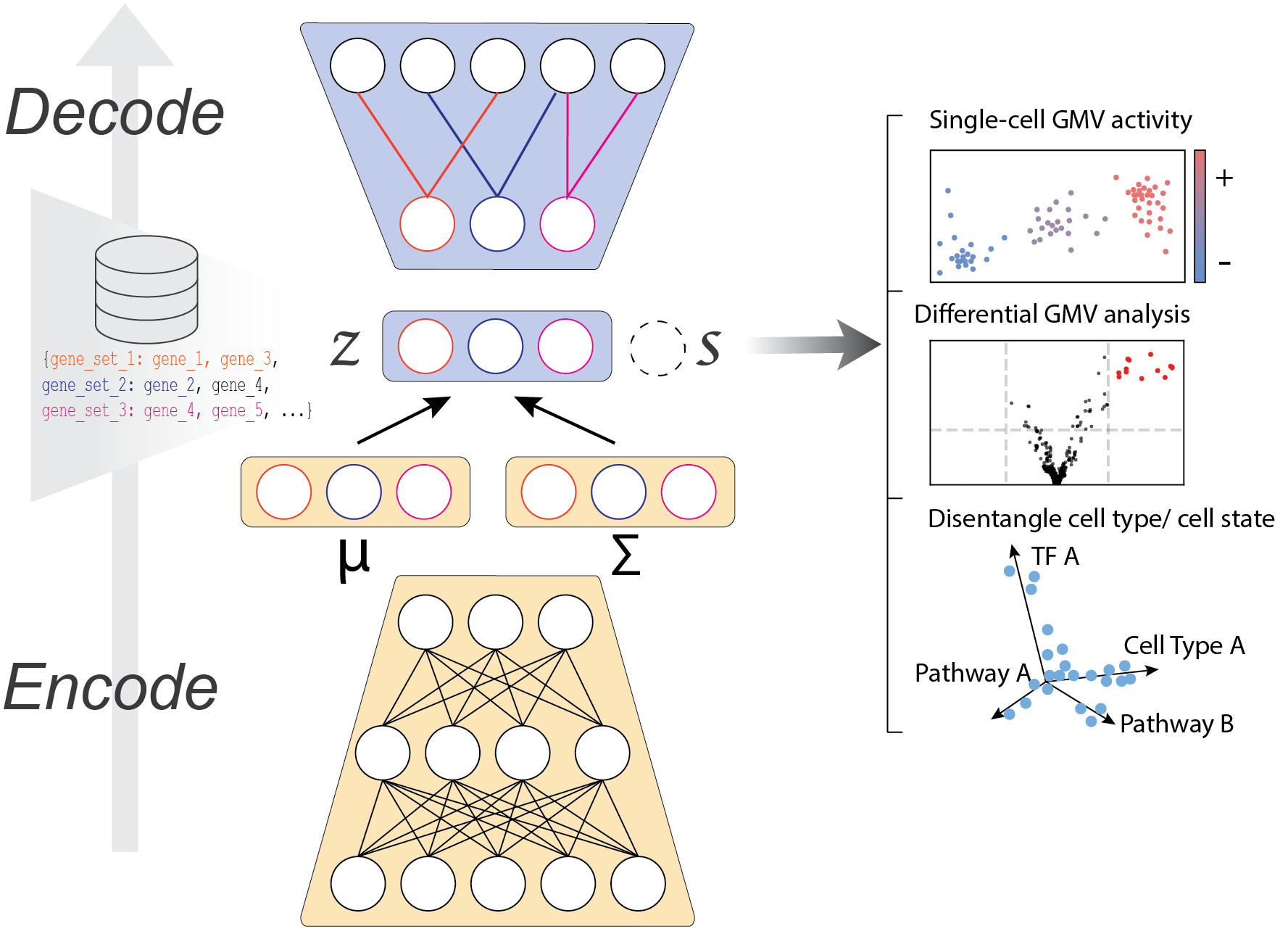
Getting started
VEGA simply requires
An Anndata single-cell dataset
GMT file(s) with the gene module membership, such as provided by databases like MSigDB
Note: We recommend to preprocess the single-cell dataset before passing it to VEGA.
Main features
VEGA provides the following features
Embed single-cell data into an interpretable latent space
Inference of gene module activities at the single-cell level
Cell type / cell state disentanglement
Alternative to enrichment methods for finding differentially activated pathways
Inspired by the differential gene expression procedure from scvi-tools, VEGA provides a Bayesian testing procedure to find significantly activated gene modules in your dataset. More information in the vega basic usage tutorial.